Hard skills are technical abilities such as market analysis, sales strategy development, and CRM skill that are essential for driving business growth.
Resume Template—Easy to Copy & Paste
Ming Huang
Minneapolis, MN 55411
(555)555-5555
Ming.Huang@example.com
Professional Summary
Dynamic Business Development Manager with 8+ years driving growth. Proven success in strategic market expansion and sales leadership. Expert in client acquisition and partnership development, achieving $5M+ in new business.
Work History
Business Development Manager
Global Solutions Inc. - Minneapolis, MN
November 2022 - November 2025
- Increased sales revenue by 20% in one year
- Developed partnerships with 15+ new clients
- Streamlined operations, boosting efficiency by 30%
Sales and Growth Leader
Innovative Ventures - Minneapolis, MN
November 2019 - October 2022
- Spearheaded market expansion increasing coverage by 40%
- Implemented CRM system, improving process by 50%
- Led team to exceed targets by 0K annually
Market Development Executive
Pioneer Tech Partners - Minneapolis, MN
November 2017 - October 2019
- Established key client relationships increasing retention by 15%
- Directed new product launch capturing 25% market share
- Coordinated cross-functional teams on projects with 2M budget
Skills
- Strategic Planning
- Market Analysis
- Sales Strategy
- Client Acquisition
- CRM Implementation
- Revenue Growth
- Team Leadership
- Partnership Development
Certifications
- Certified Business Development Expert - American Management Association
- Strategic Marketing Professional - Marketing Institute
Education
Master of Business Administration Business Management
Harvard Business School Cambridge, MA
June 2017
Bachelor of Arts Economics
University of Pennsylvania Philadelphia, PA
June 2015
Languages
- Spanish - Beginner (A1)
- French - Beginner (A1)
- German - Beginner (A1)
How to Write a Business Development Manager Resume Summary
Your resume summary is the first thing employers will notice, making it important for creating a lasting impression. As a business development manager, you should emphasize your strategic thinking and relationship-building skills to stand out. The following examples will guide you in crafting an effective summary by showcasing what resonates well with hiring managers and what to avoid:
I am an experienced business development manager with a solid background and many skills. I hope to find a position where I can use my abilities to help the company grow. A place that values teamwork and offers chances for advancement is what I’m looking for. I believe that I would be a great addition to your team if given the chance.
- Contains broad statements about experience without demonstrating specific achievements or skills
- Relies heavily on personal pronouns, making it feel less professional and more casual
- Emphasizes the applicant's desires rather than highlighting how their expertise can benefit the organization
Results-driven business development manager with 7 years of experience in driving revenue growth and market expansion for technology firms. Achieved a 30% increase in client acquisition through targeted marketing strategies and relationship management initiatives. Proficient in CRM software, data analysis, and developing strategic partnerships to improve business opportunities.
- Begins with specific years of experience highlighting a clear focus on results
- Includes measurable achievements that showcase the impact on business growth
- Lists relevant technical skills that align with industry expectations and competencies
Pro Tip
Showcasing Your Work Experience
The work experience section is important for your resume as a business development manager, serving as the main focus with the bulk of your content. Good resume templates always prioritize this section to highlight your professional journey.
Organize this part in reverse-chronological order, showcasing your previous roles and responsibilities. Use bullet points to detail your achievements and contributions in each position, making them stand out.
To further guide you, we’ll provide some examples demonstrating effective work history entries for business development managers. These examples will illustrate what works well and what should be avoided:
Business Development Manager
Hudson Corp – New York, NY
- Developed business strategies
- Worked with clients
- Attended meetings and collaborated with teams
- Conducted market research
- Lacks specific accomplishments or metrics to demonstrate success
- Too vague about client interactions and outcomes
- Focuses on general tasks rather than effective contributions
Business Development Manager
Mayflower Solutions – New York, NY
March 2020 - Current
- Develop and execute strategic plans to drive revenue growth, achieving a 30% increase in sales within the first year
- Cultivate relationships with key stakeholders, leading to the expansion of partnerships that contributed to a 25% rise in client retention rates
- Lead cross-functional teams to launch new service offerings, resulting in a successful market entry that generated $1 million in the first six months
- Starts each bullet point with action verbs that highlight the applicant's contributions
- Incorporates specific metrics and outcomes to quantify achievements effectively
- Demonstrates relevant skills such as strategic planning and relationship management integral to business development
While your resume summary and work experience are important components, don’t overlook the importance of other sections that can improve your application. To ensure every part of your resume shines, refer to our detailed guide on how to write a resume.
Top Skills to Include on Your Resume
A well-defined skills section is important for any resume, as it allows you to showcase your qualifications at a glance. This section helps potential employers quickly identify if you possess the necessary abilities for roles like business development manager.
Employers want professionals who bring together strong technical abilities and the interpersonal skills needed to collaborate and communicate effectively. Showcasing both your hard and soft skills on your resume helps demonstrate your full value as a well-rounded candidate.
Soft skills include communication, negotiation, and relationship-building which are important for fostering partnerships and ensuring collaborative success within teams.
Choosing the appropriate resume skills is important for aligning with employer expectations. Many organizations use automated screening systems to filter out applicants lacking essential criteria, so it's important to showcase relevant abilities prominently.
If you want to effectively highlight your skills, closely reviewing job postings can be beneficial. These listings often disclose the specific qualifications and experiences recruiters seek, which assists in tailoring your resume to succeed in both human evaluations and ATS screenings.
Pro Tip
10 skills that appear on successful business development manager resumes
Highlighting high-demand skills on your resume can significantly increase your chances of catching a recruiter's eye. Using resume examples can help you showcase these skills effectively, allowing you to apply with confidence and stand out in the competitive job market.
Here are 10 essential skills you should think about adding to your resume if they align with your experience and the job requirements:
Strategic planning
Sales prospecting
Negotiation tactics
Relationship management
Market analysis
Cross-functional collaboration
Project management
Lead generation
Client engagement
Data-driven decision making
Based on analysis of 5,000+ management professional resumes from 2023-2024
Resume Format Examples
Choosing the right resume format is important for a business development manager because it effectively highlights your key achievements, relevant experience, and strategic skills to potential employers.
Functional
Focuses on skills rather than previous jobs
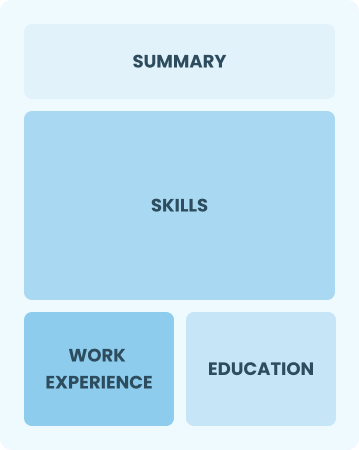
Best for:
Recent graduates and career changers with up to two years of experience
Combination
Balances skills and work history equally
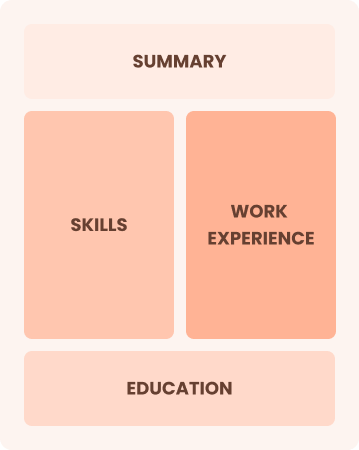
Best for:
Mid-career professionals focused on demonstrating their skills and seeking growth opportunities
Chronological
Emphasizes work history in reverse order

Best for:
Leaders driving strategic growth and innovation in business development
Frequently Asked Questions
Should I include a cover letter with my business development manager resume?
Absolutely, including a cover letter can significantly improve your application by showcasing your personality and specific qualifications for the role. It allows you to connect your experience with what the employer is seeking. For tips on crafting an effective cover letter, explore our resources on how to write a cover letter or use our Cover Letter Generator for quick assistance.
Can I use a resume if I’m applying internationally, or do I need a CV?
When applying for jobs internationally, use a CV instead of a resume as many countries require it for formal applications. To assist you, we offer resources on how to write a CV and provide helpful CV examples that align with international expectations. Explore these tools to improve your job application process.
What soft skills are important for business development managers?
Soft skills such as communication, negotiation, and interpersonal skills are essential for business development managers. These abilities enable effective collaboration with clients and colleagues, fostering trust and driving successful partnerships that contribute to business growth.
I’m transitioning from another field. How should I highlight my experience?
Highlight your transferable skills, such as negotiation, strategic thinking, and relationship-building, from previous roles. These abilities showcase your potential impact in business development, even if you lack direct experience. Share specific instances where you've driven results or fostered connections to illustrate how your background aligns with key responsibilities in this role.
Should I include a personal mission statement on my business development manager resume?
Including a personal mission statement on your resume is advisable, as it highlights your values and goals. This addition can be especially effective for companies that prioritize cultural fit or possess a clear vision. By incorporating this approach, you can resonate well with organizations focused on innovation and growth.
How do I add my resume to LinkedIn?
To increase your resume's visibility on LinkedIn, you can add your resume to LinkedIn directly or highlight critical achievements in the "About" and "Experience" sections. This approach not only showcases your qualifications but also helps recruiters easily find and connect with strong applicants like you.








