Hard skills are technical abilities such as research methodologies, time management, and understanding of academic principles that students should highlight in their studies.
Popular Student Resume Examples
Check out our top student resume examples that emphasize key skills such as time management, teamwork, and adaptability. These examples will help you effectively showcase your academic achievements and extracurricular activities.
Ready to build your standout resume? Our Resume Builder offers user-friendly templates designed specifically for students to help you shine in your job applications.
Recommended
Student worker resume
The resume's clean formatting and professional resume fonts selection improve readability, helping this early-career professional present their qualifications clearly. This attention to detail shows their organizational skills and commitment to quality, making a positive impression on potential employers.
Campus ambassador resume
This resume effectively balances skills such as public relations and event planning with hands-on experience in various roles. This combination allows employers to see how the applicant has used these skills to achieve tangible results, improving their understanding of the applicant's potential contributions in a professional setting.
Resident assistant resume
This resume effectively uses bullet points to clearly present diverse experiences, making it easy for hiring managers to quickly assess qualifications. The thoughtful use of spacing and categories ensures that key accomplishments stand out, providing a comprehensive overview without overwhelming the reader.
Resume Template—Easy to Copy & Paste
Suki Smith
Riverside, CA 92512
(555)555-5555
Suki.Smith@example.com
Professional Summary
Innovative student with 9 years of experience in data science, analytics, and predictive modeling. Expertise in Python, machine learning, and big data, driving impact through 25% operational improvements and exceptional project outcomes.
Work History
Student
AI Learning Center - Riverside, CA
January 2024 - October 2025
- Achieved 95% course completion rate in AI program
- Participated in 10+ collaborative coding bootcamps
- Completed research project leading to 25% system improvement
Data Analyst
Core Analytics Hub - San Francisco, CA
January 2020 - December 2023
- Improved data accuracy by over 30% using new validation tools
- Designed dashboards utilized by 75+ stakeholders
- Analyzed 10TB of financial data, optimizing insights by 40%
Junior Business Analyst
Precision Analytics Inc. - Riverside, CA
January 2016 - December 2019
- Streamlined reporting reducing errors by 25%
- Managed data for 2M+ client projects efficiently
- Provided actionable recommendations improving ROI by 15%
Languages
- Spanish - Beginner (A1)
- German - Intermediate (B1)
- Mandarin - Beginner (A1)
Skills
- Data Analysis
- Machine Learning
- Statistical Modeling
- Python Programming
- Visualization Tools
- Big Data Processing
- Database Management
- AI Research
Certifications
- Certified Data Analyst - International Data Certification Board
- Google Cloud Professional Data Engineer - Google Cloud
- Advanced Data Analytics Certification - Microsoft Training Academy
Education
Master's Data Science
University of California Berkeley Berkeley, California
December 2015
Bachelor's Computer Science
University of Washington Seattle, Washington
June 2014
How to Write a Student Resume Summary
Your resume summary is the first impression you make on hiring managers, so it’s important to present your qualifications effectively. This section should reflect your strengths and relevant experiences as a student, giving employers insight into your potential contributions.
As a student, you’ll want to highlight skills such as teamwork, communication, and problem-solving acquired through coursework or extracurricular activities. Showcasing these attributes can set you apart from other applicants.
To better understand what makes an powerful resume summary, take a look at the examples provided below. They will illustrate effective strategies and common pitfalls:
I am a college student with some experience in various projects and activities. I want to find an internship that will allow me to grow my skills and gain new experiences. A place where I can learn and develop professionally would be great. I believe I can be a valuable addition if given the chance.
- Lacks specific details about skills or relevant experiences, making it hard for employers to gauge the job seeker's qualifications
- Focuses too much on personal aspirations rather than how the applicant can contribute to the organization
- Uses generic phrases that do not highlight unique strengths or achievements, leading to a bland impression
Motivated student with 2 years of experience in volunteer tutoring, specializing in mathematics and science for high school students. Improved student performance by 20% on average through personalized study plans and engaging learning activities. Proficient in Microsoft Office Suite and educational technology tools, with strong communication and teamwork skills.
- Highlights specific experience level relevant to the education field
- Includes a quantifiable achievement that showcases effectiveness in improving student outcomes
- Mentions technical skills and competencies that are valuable for teaching and collaboration
Pro Tip
Showcasing Your Work Experience
The work experience section is important for your resume as a student, where you'll provide the bulk of your content. Good resume templates always feature this section prominently to highlight relevant experiences.
You should organize this part chronologically, listing your roles in reverse order. Use bullet points to showcase key achievements and responsibilities that demonstrate your skills and contributions during your studies or internships.
Now, let’s look at a couple of examples that illustrate effective entries for students' work experience. These will help you understand what works well and what to avoid.
Student
Governor High School – Detroit, MI
- Attended classes regularly
- Worked on group projects
- Participated in extracurricular activities
- Studied for exams and quizzes
- No specifics about subjects or achievements
- Bullet points are vague and do not highlight skills or contributions
- Lacks emphasis on leadership roles or measurable successes
Student
Arizona State University – Tempe, AZ
September 2021 - Present
- Maintain a GPA of 3.8 while balancing coursework and part-time employment, demonstrating time management skills
- Lead a team project to develop a mobile app that earned recognition at the university's annual tech fair for innovation
- Volunteer as a tutor for underclassmen in mathematics, helping increase their understanding and performance in the subject
- Uses strong action verbs to clearly convey accomplishments and responsibilities
- Incorporates specific metrics like GPA and project recognition to highlight performance
- Demonstrates relevant skills such as leadership and mentoring within academic settings
While your resume summary and work experience are important components, it's essential not to overlook the other sections that can improve your application. For more detailed guidance on crafting a standout resume, be sure to explore our comprehensive guide on how to write a resume.
Top Skills to Include on Your Resume
A skills section is an important part of your resume as it allows you to showcase your qualifications at a glance. It helps potential employers quickly identify if you have the abilities needed for their position.
As a student, you want to showcase your full range of abilities. Including a mix of hard and soft skills is essential to showing employers you have the required subject matter expertise and interpersonal abilities to thrive in the role.
Soft skills, on the other hand, are interpersonal qualities like adaptability, teamwork, and effective communication that matter for collaboration with peers and fostering a positive learning environment.
When selecting your resume skills, it's important to align them with what employers expect. Many organizations use automated systems that filter out applicants lacking essential skills for the position.
To improve your chances of standing out, carefully examine job postings for specific skills mentioned. This practice not only helps you tailor your resume skills for recruiters but also ensures compatibility with ATS systems that scan for these qualifications.
Pro Tip
10 skills that appear on successful student resumes
Highlighting essential skills on your resume can significantly increase your chances of catching a recruiter's eye. You can see these high-demand skills reflected in our resume examples, making it easier for you to present yourself confidently.
By the way, here are 10 key skills you might consider adding to your resume if they align with your experience and job requirements:
Fast learner
Positive attitude
Attention to detail
Flexible schedule
Team collaboration
Active listening
Time management
Energetic and enthusiastic
Basic housekeeping
Punctual and reliable
Based on analysis of 5,000+ education professional resumes from 2023-2024
Resume Format Examples
Choosing the appropriate resume format is important because it highlights your most relevant skills and experiences, enabling potential employers to quickly assess your growth and suitability for the role.
Functional
Focuses on skills rather than previous jobs
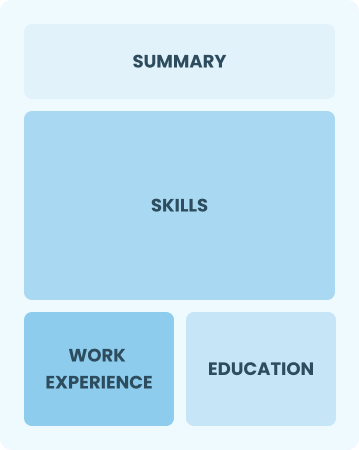
Best for:
Recent grads and career changers with up to two years' experience
Combination
Balances skills and work history equally
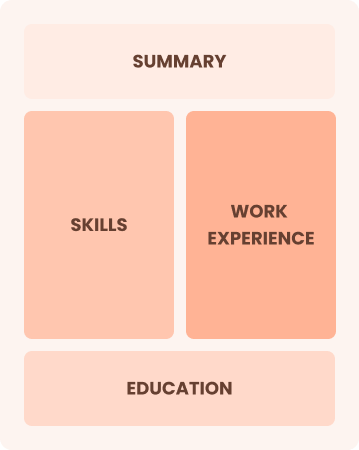
Best for:
Mid-career professionals eager to demonstrate their skills and pursue new opportunities
Chronological
Emphasizes work history in reverse order

Best for:
Accomplished students excelling in advanced studies with strong subject-matter expertise
Frequently Asked Questions
Should I include a cover letter with my student resume?
Absolutely. Including a cover letter can significantly improve your application by showcasing your personality and detailing your qualifications. It gives you the chance to connect directly with hiring managers. If you need assistance, check out our comprehensive guide on how to write a cover letter for crafting a standout application or use our Cover Letter Generator for quick support.
Can I use a resume if I’m applying internationally, or do I need a CV?
When applying for jobs abroad, use a CV instead of a resume as it provides a comprehensive overview of your academic background and work experiences. Explore our resources on how to write a CV to learn the essential formatting techniques and content guidelines that align with international expectations. Additionally, check out CV examples to see how these principles are applied in real-world scenarios.
What soft skills are important for students?
Soft skills like communication, collaboration, and time management are essential for students. These interpersonal skills enable effective teamwork and strong relationships with peers and instructors, fostering a positive learning environment that improves academic success.
I’m transitioning from another field. How should I highlight my experience?
Highlight transferable skills gained from previous roles, such as communication, teamwork, and adaptability. These abilities show your potential to thrive in a new environment, even if you lack direct experience. Provide concrete examples that connect your past successes to the key responsibilities of the student role, demonstrating your readiness to excel.
Where can I find inspiration for writing my cover letter as a student?
For those applying for student positions, cover letter examples can be invaluable. These samples provide inspiration for content ideas, help with formatting, and showcase how to effectively present your qualifications. Referring to them can improve the quality of your application materials and increase your chances of standing out to employers.
How do I write a resume with no experience?
When applying for student positions with limited work experience, consider using a resume with no experience format to highlight relevant coursework, group projects, and extracurricular activities. Showcase skills like teamwork, adaptability, and time management. Remember, employers value your potential and enthusiasm to learn. By focusing on your passion for the field, you’ll stand out even without extensive experience.








