Hard skills are technical abilities, including data analysis, experimental design, and skill in laboratory techniques that researchers must master to conduct effective studies.
Popular Researcher Resume Examples
Check out our top researcher resume examples that emphasize critical skills such as data analysis, problem-solving, and innovative thinking. These examples demonstrate how to effectively present your contributions in research-driven environments.
Need a polished resume? Our Resume Builder offers customizable templates specifically designed for researchers to help you shine in your career pursuit.
Recommended
Entry-level researcher resume
This entry-level resume highlights their achievements in improving project outcomes and publishing high-impact papers, showcasing strong analytical skills and leadership. New researchers should focus on demonstrating their technical proficiencies and ability to contribute effectively to team projects despite limited industry experience.
Mid-career researcher resume
This resume effectively showcases the job seeker’s qualifications by emphasizing leadership in research and significant achievements. The structured presentation of past roles and accomplishments clearly indicates readiness for advanced responsibilities, demonstrating a solid trajectory in the research field.
Experienced researcher resume
The work experience section illustrates the applicant's extensive research background and data analysis skills, notably analyzing over 50 datasets and publishing 12 articles annually. The clear formatting allows hiring managers to quickly assess achievements, making it ideal for applicants in competitive fields.
Resume Template—Easy to Copy & Paste
Daniel Lee
Indianapolis, IN 46201
(555)555-5555
Daniel.Lee@example.com
Professional Summary
Innovative researcher with 8 years in AI, data analysis, and tech innovation. Proven track record of enhancing project efficiency by over 30% using scientific methods and advanced data modeling. Adept at leading projects and publishing impactful research.
Work History
Researcher
Innovate Research Center - Indianapolis, IN
May 2022 - December 2025
- Led 5 projects, increasing efficiency by 30%
- Collaborated with 4 teams to develop AI models
- Published 12 papers in peer-reviewed journals
Research Analyst
Quantum Solutions - Indianapolis, IN
April 2018 - April 2022
- Analyzed data, improving accuracy by 20%
- Presented findings to 10 industry conferences
- Developed tools, saving 200 hours annually
Research Assistant
Future Insights Lab - Greenfield, IN
April 2015 - April 2018
- Supported 8 research projects with data collection
- Managed datasets, enhancing processing speed by 25%
- Authored 5 technical reports for publication
Languages
- Spanish - Beginner (A1)
- French - Intermediate (B1)
- Mandarin - Beginner (A1)
Skills
- Data Analysis
- Quantitative Research
- Machine Learning
- Statistical Modeling
- Project Management
- Data Visualization
- Scientific Writing
- Problem Solving
Certifications
- Certified Data Scientist - Data Science Council of America
- AI Product Manager - AI Academy
- Advanced R Programming - Coursera
Education
Master of Science Artificial Intelligence
Massachusetts Institute of Technology Cambridge, MA
June 2015
Bachelor of Science Computer Science
University of California, Berkeley Berkeley, CA
June 2013
How to Write a Researcher Resume Summary
Your resume summary is the first opportunity to capture an employer's attention, making it important to present a compelling introduction. As a researcher, you should emphasize your analytical skills, relevant findings, and contributions to the field that set you apart from other job seekers.
To guide you in crafting an effective summary, review the following examples that illustrate successful approaches and common pitfalls:
I am a dedicated researcher with a wide range of experience in various fields. I hope to find a job that allows me to use my skills and knowledge effectively. A company that values innovation and offers chances for advancement would be perfect for me. I believe I can contribute positively if given the chance.
- Contains general statements about the applicant's experience without specific examples or achievements
- Overuses personal language, making it feel less professional and more like filler content
- Emphasizes what the job seeker is looking for in a role rather than highlighting what they can bring to the organization
Results-driven researcher with over 6 years of experience in clinical trials and data analysis, focusing on oncology studies. Successfully improved patient recruitment rates by 25% through targeted outreach strategies and refined eligibility screening processes. Proficient in statistical software such as SPSS and SAS, as well as managing cross-functional teams to ensure compliance with regulatory standards.
- Starts with specific experience duration and research focus area
- Highlights quantifiable achievement that showcases measurable impact on research efficiency
- Includes relevant technical skills that align with industry expectations for researchers
Pro Tip
For inspiration, there are plenty of resume objective examples available tailored to your field.
Showcasing Your Work Experience
The work experience section is important for your resume as a researcher, containing the bulk of your content. Good resume templates always emphasize this section to highlight your relevant experience.
This area should be organized in reverse-chronological order, detailing your previous positions. Use bullet points to succinctly describe your achievements and contributions in each research role.
To illustrate effective entries, we will present a couple of examples that demonstrate what works well and what pitfalls to avoid:
Researcher
Global Research Institute – New York, NY
- Conducted research activities.
- Collected and analyzed data.
- Collaborated with team members.
- Assisted in preparing reports.
- Lacks specific details about the types of research conducted
- Bullet points do not showcase any measurable outcomes or impacts
- Focuses on generic tasks rather than highlighting unique contributions or skills
Researcher
Innovative Solutions Lab – San Francisco, CA
March 2020 - Current
- Conduct in-depth analysis of data trends to support the development of new product innovations, increasing project efficiency by 30%.
- Collaborate with cross-functional teams to design and execute experiments, achieving a 15% increase in accuracy of predictive models.
- Mentor junior researchers through structured training sessions, fostering a collaborative environment that improves team productivity.
- Each bullet point starts with strong action verbs that clearly outline the applicant's contributions.
- Incorporates specific metrics that quantify achievements, showing tangible results of efforts
- Highlights relevant skills and experiences that align with industry expectations for researchers
While your resume summary and work experience are important components, don’t overlook the importance of other sections. Each part plays a role in presenting your qualifications effectively. For more insights on crafting a standout resume, be sure to explore our complete guide on how to write a resume.
Top Skills to Include on Your Resume
A skills section is an important component of your resume, as it allows you to highlight your qualifications at a glance. This section can help potential employers quickly identify whether you possess the necessary abilities for the researcher role.
Every resume should have a mix of hard skills and soft skills. Including a diverse range of relevant skills will make your application much stronger.
Soft skills such as critical thinking, collaboration, and adaptability are essential for fostering innovative solutions and working efficiently within diverse teams.
When selecting skills for your resume, it's important to align with what employers expect from applicants. Many organizations use automated systems to filter out applicants lacking essential resume skills, making it important to tailor your skills accordingly.
To effectively prioritize your skills, take the time to review job postings relevant to your field. These listings provide valuable insights into which specific abilities recruiters and ATS systems are seeking, allowing you to improve your chances of standing out in the application process.
Pro Tip
10 skills that appear on successful researcher resumes
Improving your resume by adding sought-after skills can significantly attract the attention of recruiters for researcher roles. You can find these skills highlighted in our resume examples, enabling you to apply confidently and effectively.
By the way, consider incorporating relevant skills from this selection into your resume if they align with your qualifications and job expectations:
Analytical thinking
Attention to detail
Data analysis
Technical writing
Project management
Statistical software skill
Collaboration
Problem-solving
Time management
Research methodology
Based on analysis of 5,000+ sciences professional resumes from 2023-2024
Resume Format Examples
Selecting the appropriate resume format is important since it highlights your most relevant research skills, experience, and career development in a clear and effective way.
Functional
Focuses on skills rather than previous jobs
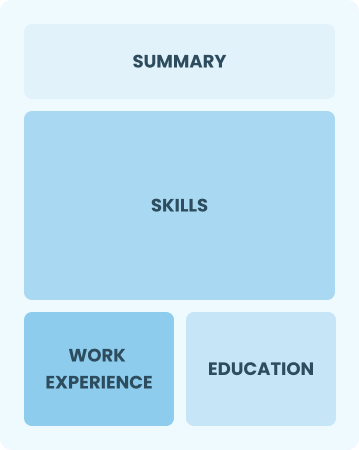
Best for:
Recent graduates and career changers with up to two years of experience
Combination
Balances skills and work history equally
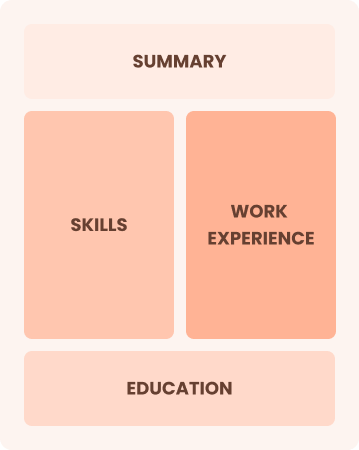
Best for:
Mid-career professionals eager to highlight their skills and growth potential
Chronological
Emphasizes work history in reverse order

Best for:
Seasoned professionals leading innovative research initiatives
Researcher Salaries in the Highest-Paid States
Our researcher salary data is based on figures from the U.S. Bureau of Labor Statistics (BLS), the authoritative source for employment trends and wage information nationwide.
Whether you're entering the workforce or considering a move to a new city or state, this data can help you gauge what fair compensation looks like for researchers in your desired area.
Frequently Asked Questions
Should I include a cover letter with my researcher resume?
Absolutely, including a cover letter can significantly improve your application by allowing you to showcase your personality and clarify your qualifications. It gives you the chance to connect with hiring managers on a deeper level.
For tips on crafting a powerful cover letter, consider checking out our comprehensive guide on how to write a cover letter or use our easy-to-navigate Cover Letter Generator to get started quickly.
Can I use a resume if I’m applying internationally, or do I need a CV?
When applying for jobs outside the U.S., a CV is often required instead of a resume. A CV offers a comprehensive view of your academic background and professional experience. For guidance on formatting, check out how to write a CV with detailed tips. Additionally, explore our CV examples tailored for international applications.
What soft skills are important for researchers?
Soft skills such as critical thinking, collaboration, and effective communication are essential for researchers. These interpersonal skills foster teamwork and facilitate the sharing of ideas, leading to innovative solutions and stronger partnerships in research projects.
I’m transitioning from another field. How should I highlight my experience?
Highlight your transferable skills such as communication, analytical thinking, and teamwork from prior roles. These abilities illustrate your potential to add value in research positions, even if you lack direct experience.
By sharing concrete examples that relate your previous successes to the responsibilities of a researcher, you can effectively showcase how you can make a meaningful impact.
Should I use a cover letter template?
Yes, using a cover letter template tailored for researchers can improve your application by providing a clear structure and helping you highlight critical skills such as data analysis, experimental design, and publication success that are essential to impress hiring managers.
How do I add my resume to LinkedIn?
To add your resume to LinkedIn and improve its visibility, upload it directly or highlight essential achievements in the "About" and "Experience" sections. This approach allows recruiters and hiring managers to easily find qualified researchers like you, increasing your chances of being noticed.








